Protein Domains
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What are protein domains?
Domains are highly conserved units within a protein that normally have a specific structure and function. These units are the "building blocks" of a protein, and they usually contribute to the overall functions and interactions of the protein. [1] Domains have a three-dimensional structure and can often exist independently. Because they are so highly conserved, domains are used and re-used in a wide variety of proteins. [2]
Domains in XPA
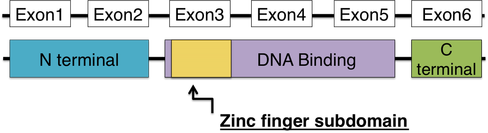
XPA is a small and simple protein; at 273 amino acids, there isn't a lot of room for complex domain structure. The XPA protein has an N-terminal domain, a C-terminal domain, and a DNA binding domain with a zinc finger subdomain. The N-terminal domain is responsible for producing enzymes needed for nucleotide excision repair, and the C-terminal domain has not been well characterized. [3,4] However, of the three domains, the DNA binding domain is the most important. The DNA binding domain allows XPA to bind to damaged DNA, and it does this with the help of the zinc finger subdomain. [4] Zinc fingers are well conserved in nature and can be found in a wide variety of proteins that involve DNA binding. Mutations that fall within the DNA binding domain tend to be the most severe. [5]
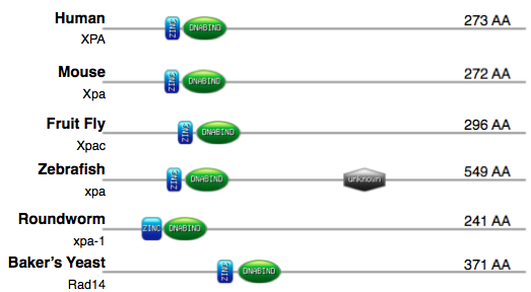
Figure 2 was created with My Domains Image Creator from Prosite, and relative domain locations were obtained using InterPro. InterPro and databases of the like provide functional analysis of protein sequences by classifying and identifying domains and other important sites. [6] With the help of these databases, domains and other functional elements can be compared across a variety of organisms.
Discussion
The figure above shows the relative location of each domain and the size of the protein across model organisms. It is easy to see that XPA is relatively well conserved across organisms, especially the position of the zinc finger subdomain and DNA binding domain. The close proximity of these two domains in all model organisms reinforces the importance that the domains work together for proper function. One notable difference can be found in zebrafish; zebrafish have a surprisingly large protein that contains an extra domain. This domain's function is unknown, but the domain is commonly found in eukaryotes and contains a downstream promoter element (DPE) sequence motif. [7] A DPE is a type of promoter that helps initiate transcription, thus, this possible promoter within the protein may suggest additional functions of XPA in zebrafish.
The figure above shows the relative location of each domain and the size of the protein across model organisms. It is easy to see that XPA is relatively well conserved across organisms, especially the position of the zinc finger subdomain and DNA binding domain. The close proximity of these two domains in all model organisms reinforces the importance that the domains work together for proper function. One notable difference can be found in zebrafish; zebrafish have a surprisingly large protein that contains an extra domain. This domain's function is unknown, but the domain is commonly found in eukaryotes and contains a downstream promoter element (DPE) sequence motif. [7] A DPE is a type of promoter that helps initiate transcription, thus, this possible promoter within the protein may suggest additional functions of XPA in zebrafish.
References
1. "Introduction to Protein Classification at the EBI "What Are Protein Domains?" What Are Protein Domains? European Bioinformatics Institute, 2012. http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains. Retrieved May 05, 2014.
2. Feldman, Howard J. (2012) Identifying structural domains of proteins using clustering. BMC Bioinformatics, 13:286. doi:10.1186/1471-2105-13-286 http://www.biomedcentral.com/content/pdf/1471-2105-13-286.pdf
3. Bartels, Claudine L., Lambert, Muriel W. (2007) Domains in the XPA protein important in its role as a processivity factor. Biochemical and Biophysical Research Communications, 356, 219–225. Available from http://www.ncbi.nlm.nih.gov/pubmed/17349973.
4. InterPro. http://www.ebi.ac.uk/interpro/entry/IPR022656. Retrieved April 30, 2014.
5. Amr, Khalda, Messaoud, Olfa, El Darouti, Mohamad, Abdelha, Sonia, El-Kamah, Ghada. (2014) Mutational specturm of Xeroderma pigmentosum group A in Egyptian patients. Gene, 533, 52-56. Available from http://www.ncbi.nlm.nih.gov/pubmed/24135642.
6. Sarah Hunter; Philip Jones; Alex Mitchell; Rolf Apweiler; Teresa K. Attwood; Alex Bateman; Thomas Bernard; David Binns; Peer Bork; Sarah Burge; Edouard de Castro; Penny Coggill; Matthew Corbett; Ujjwal Das; Louise Daugherty; Lauranne Duquenne; Robert D. Finn; Matthew Fraser; Julian Gough; Daniel Haft; Nicolas Hulo; Daniel Kahn; Elizabeth Kelly; Ivica Letunic; David Lonsdale; Rodrigo Lopez; Martin Madera; John Maslen; Craig McAnulla; Jennifer McDowall; Conor McMenamin; Huaiyu Mi; Prudence Mutowo-Muellenet; Nicola Mulder; Darren Natale; Christine Orengo; Sebastien Pesseat; Marco Punta; Antony F. Quinn; Catherine Rivoire; Amaia Sangrador-Vegas; Jeremy D. Selengut; Christian J. A. Sigrist; Maxim Scheremetjew; John Tate; Manjulapramila Thimmajanarthanan; Paul D. Thomas; Cathy H. Wu; Corin Yeats; Siew-Yit Yong (2011). InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Research 2011; doi: 10.1093/nar/gkr948
7. Family: DUF4585 (PF15232). Pfam. http://pfam.xfam.org/family/PF15232. Retrieved May 10, 2014.
1. "Introduction to Protein Classification at the EBI "What Are Protein Domains?" What Are Protein Domains? European Bioinformatics Institute, 2012. http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains. Retrieved May 05, 2014.
2. Feldman, Howard J. (2012) Identifying structural domains of proteins using clustering. BMC Bioinformatics, 13:286. doi:10.1186/1471-2105-13-286 http://www.biomedcentral.com/content/pdf/1471-2105-13-286.pdf
3. Bartels, Claudine L., Lambert, Muriel W. (2007) Domains in the XPA protein important in its role as a processivity factor. Biochemical and Biophysical Research Communications, 356, 219–225. Available from http://www.ncbi.nlm.nih.gov/pubmed/17349973.
4. InterPro. http://www.ebi.ac.uk/interpro/entry/IPR022656. Retrieved April 30, 2014.
5. Amr, Khalda, Messaoud, Olfa, El Darouti, Mohamad, Abdelha, Sonia, El-Kamah, Ghada. (2014) Mutational specturm of Xeroderma pigmentosum group A in Egyptian patients. Gene, 533, 52-56. Available from http://www.ncbi.nlm.nih.gov/pubmed/24135642.
6. Sarah Hunter; Philip Jones; Alex Mitchell; Rolf Apweiler; Teresa K. Attwood; Alex Bateman; Thomas Bernard; David Binns; Peer Bork; Sarah Burge; Edouard de Castro; Penny Coggill; Matthew Corbett; Ujjwal Das; Louise Daugherty; Lauranne Duquenne; Robert D. Finn; Matthew Fraser; Julian Gough; Daniel Haft; Nicolas Hulo; Daniel Kahn; Elizabeth Kelly; Ivica Letunic; David Lonsdale; Rodrigo Lopez; Martin Madera; John Maslen; Craig McAnulla; Jennifer McDowall; Conor McMenamin; Huaiyu Mi; Prudence Mutowo-Muellenet; Nicola Mulder; Darren Natale; Christine Orengo; Sebastien Pesseat; Marco Punta; Antony F. Quinn; Catherine Rivoire; Amaia Sangrador-Vegas; Jeremy D. Selengut; Christian J. A. Sigrist; Maxim Scheremetjew; John Tate; Manjulapramila Thimmajanarthanan; Paul D. Thomas; Cathy H. Wu; Corin Yeats; Siew-Yit Yong (2011). InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Research 2011; doi: 10.1093/nar/gkr948
7. Family: DUF4585 (PF15232). Pfam. http://pfam.xfam.org/family/PF15232. Retrieved May 10, 2014.
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison