Gene Ontology
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What is Gene Ontology?
The Gene Ontology project is a collaborative effort to classify a wide variety of gene products across a multitude of model organisms. [1] This project originated due to the lack of a common language to describe how the cell functions. Without equivalent terms to describe the various cell processes, it was incredibly difficult to compare these processes across organisms.
In order to accurately define the function or location of a certain gene product, three terms were developed. A gene's ontology is defined by its cellular component, biological process, and molecular function. [1] Cellular component describes the component of the cell that the gene localizes, either in a gene product group or an organelle. The biological process is a series of events that occur as a result of the gene, and is generally defined by having multiple steps. Molecular function describes the activities of the gene at a molecular level; it specifically defines the activity that is occurring. [1]
Ontology of the XPA gene
The results from the Gene Ontology database showed that XPA has six cellular components, eight biological processes, and five molecular functions:
The results from the Gene Ontology database showed that XPA has six cellular components, eight biological processes, and five molecular functions:
|
Cellular Component
Cytoplasm (GO:0005737) Golgi apparatus (GO:0005794) Intercellular bridge (GO:0045171) Nucleolus (GO:0005730) Nucleoplasm (GO:0005654) Nucleus (GO:0005634) |
Biological Process
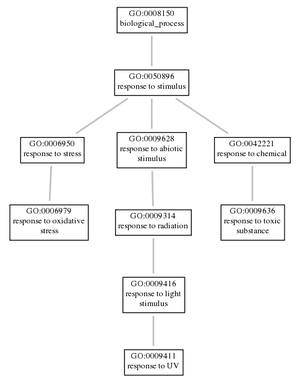
DNA repair (GO:0006281) Intrinsic apoptotic signaling pathway in response to DNA damage (GO:0008630) Multicellular organism growth (GO:0035264) Nucleotide-excision repair (GO:0006289) Nucleotide-excision repair, DNA damage removal (GO:0000718) Response to oxidative stress (GO:0006979) Response to toxic substance (GO:0009636) Response to UV (GO:0009411) |
Molecular Function
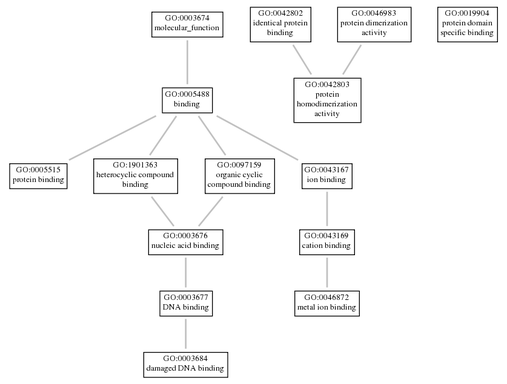
Damaged DNA binding (GO:0003684) Metal ion binding (GO:0046872) Protein binding (GO:0005515) Protein domain specific binding (GO:0019904) Protein homodimerization activity (GO:0042803) |
As shown by the above results, the XPA gene is an important factor in many biological processes, molecular functions, and cellular components. Gene ontology can be studied by observing the connections that a process, function, or component has with has with other factors within the same category. However, if all factors in one category of ontology were observed at once for XPA, the results would be overwhelming and difficult to interpret. Thus, a few related biological processes and the molecular functions are shown in two different figures below. Depending on the complexity for the particular gene, flowcharts can incorporate all ontologies of a category, or might use just a few.
Discussion
The XPA gene is part of nucleotide excision repair, a very complex DNA repair process, and thus it makes sense is involved within many systems in the cell. Upon analyzing the various molecular functions, biological processes, and cellular components, XPA is present and active across the entire cell. This is not surprising, because DNA damage is not necessarily contained to one specific part of the cell. Another ontology that fit my expectations was the molecular function. Nucleotide excision repair operates as a complex of proteins, which requires a significant amount of binding in order for the proteins to communicate with each other. Almost every molecular function for XPA involves binding, and the related molecular functions in Figure 2 incorporate binding as well. Although XPA doesn't perform one function or process entirely on its own, it is a vital addition to the other proteins involved.
The XPA gene is part of nucleotide excision repair, a very complex DNA repair process, and thus it makes sense is involved within many systems in the cell. Upon analyzing the various molecular functions, biological processes, and cellular components, XPA is present and active across the entire cell. This is not surprising, because DNA damage is not necessarily contained to one specific part of the cell. Another ontology that fit my expectations was the molecular function. Nucleotide excision repair operates as a complex of proteins, which requires a significant amount of binding in order for the proteins to communicate with each other. Almost every molecular function for XPA involves binding, and the related molecular functions in Figure 2 incorporate binding as well. Although XPA doesn't perform one function or process entirely on its own, it is a vital addition to the other proteins involved.
References
1. The Gene Ontology Consortium. "Gene ontology: tool for the unification of biology." Nature Genetics. May 2000; 25(1):25-9. Available from http://www.ncbi.nlm.nih.gov/pubmed/10802651
1. The Gene Ontology Consortium. "Gene ontology: tool for the unification of biology." Nature Genetics. May 2000; 25(1):25-9. Available from http://www.ncbi.nlm.nih.gov/pubmed/10802651
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison