Protein Interactions
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What are protein interactions?
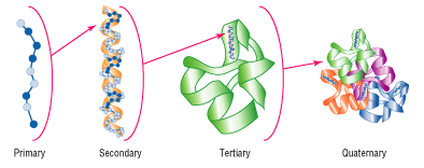 Figure 1: Proteins are composed of four structural levels.
Figure 1: Proteins are composed of four structural levels.
For some proteins, proper functioning is dependent on the interactions with neighboring proteins. Protein interactions are when a protein binds with two or more neighboring proteins in order to carry out their biological function. [1] The interaction of one protein with one or more other proteins makes up its quaternary structure, as shown in the figure. Proteins are not always expressed in the same way across varying biological conditions, however. For this reason, it is important to study the interactions between other proteins, as the function of the binding partners can give insight to the protein of interest. [2] The STRING database can generate protein interaction networks of a given protein of interest.
XPA in STRING
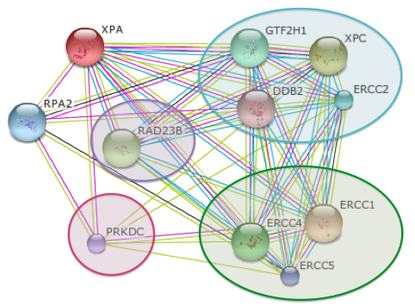
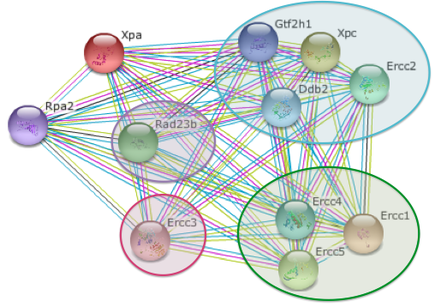
The XPA protein and its binding partners are incredibly well studied, as is shown in the two figures below. Each colored line has a meaning in the STRING database. The pink line indicates that an experiment was performed for that interaction; the light green line indicates that the proteins have been referenced together in the literature; the light blue line indicates that the proteins are coordinated in the same data set within a database; and the black line indicates that the referenced proteins are expressed at the same time.
The XPA protein and its binding partners are incredibly well studied, as is shown in the two figures below. Each colored line has a meaning in the STRING database. The pink line indicates that an experiment was performed for that interaction; the light green line indicates that the proteins have been referenced together in the literature; the light blue line indicates that the proteins are coordinated in the same data set within a database; and the black line indicates that the referenced proteins are expressed at the same time.
The "blue group" represents the proteins that bind in nucleotide excision repair early on in the process.
The "green group" represents the proteins that bind in nucleotide excision repair late in the process.
The protein in purple is a yeast homolog found in both humans and mice; not part of the nucleotide excision repair protein complex.
The protein in pink highlights two different proteins that are either found in mice or humans, but not both.
The "green group" represents the proteins that bind in nucleotide excision repair late in the process.
The protein in purple is a yeast homolog found in both humans and mice; not part of the nucleotide excision repair protein complex.
The protein in pink highlights two different proteins that are either found in mice or humans, but not both.
Discussion
Upon analyzing the STRING networks for human and mouse, a few interesting patterns surfaced. Humans and mice are closely related, so it came as no surprise that the interacting proteins in DNA repair are almost identical. There are two proteins that differ between the two organisms, however. In humans, the unique protein is PRKDC, otherwise known as DNA-dependent protein kinase catalytic subunit. This kinase acts as a molecular sensor for DNA damage and aids in the physical breakage of DNA that is necessary for repair. [3] This function does not come as a surprise, but it is interesting that this seemingly important kinase is not found in mice. Conversely, the protein Ercc3 is found in mice but not in humans. According to STRING, Ercc3 is an excision repair cross-complementing rodent repair deficiency, complementation group 3 gene; this essentially means that the protein is involved in nucleotide excision repair in mice. In humans, however, ERCC3 is a gene that codes for the production of the XPB protein. [4] XPB binds to another DNA repair protein called XPD, and these two components make up a transcription factor called general transcription factor IIH (TFIIH). [5] This is interesting because the XPD protein is also known as ERCC2, and this protein is present in both the human and mouse interaction network; however, its binding partner, XPB, is not. [6] Further research should be performed to investigate these discrepancies.
One other protein of interest, which is shared by both organisms, is RAD23b. The protein is found in both organisms, but according to STRING, is actually a yeast homolog. The protein plays a central role in degradation of misfolded proteins and in DNA repair. The protein also stabilizes the protein XPC, which is also included in both networks. This stabilization is vital because XPC is the first protein to recognize DNA distortion. [5] Overall, the protein interaction networks were not surprising. Nucleotide excision repair works as a large complex of proteins, so it makes sense that almost all of the proteins interact with each other in some way.
Upon analyzing the STRING networks for human and mouse, a few interesting patterns surfaced. Humans and mice are closely related, so it came as no surprise that the interacting proteins in DNA repair are almost identical. There are two proteins that differ between the two organisms, however. In humans, the unique protein is PRKDC, otherwise known as DNA-dependent protein kinase catalytic subunit. This kinase acts as a molecular sensor for DNA damage and aids in the physical breakage of DNA that is necessary for repair. [3] This function does not come as a surprise, but it is interesting that this seemingly important kinase is not found in mice. Conversely, the protein Ercc3 is found in mice but not in humans. According to STRING, Ercc3 is an excision repair cross-complementing rodent repair deficiency, complementation group 3 gene; this essentially means that the protein is involved in nucleotide excision repair in mice. In humans, however, ERCC3 is a gene that codes for the production of the XPB protein. [4] XPB binds to another DNA repair protein called XPD, and these two components make up a transcription factor called general transcription factor IIH (TFIIH). [5] This is interesting because the XPD protein is also known as ERCC2, and this protein is present in both the human and mouse interaction network; however, its binding partner, XPB, is not. [6] Further research should be performed to investigate these discrepancies.
One other protein of interest, which is shared by both organisms, is RAD23b. The protein is found in both organisms, but according to STRING, is actually a yeast homolog. The protein plays a central role in degradation of misfolded proteins and in DNA repair. The protein also stabilizes the protein XPC, which is also included in both networks. This stabilization is vital because XPC is the first protein to recognize DNA distortion. [5] Overall, the protein interaction networks were not surprising. Nucleotide excision repair works as a large complex of proteins, so it makes sense that almost all of the proteins interact with each other in some way.
References
1. EMBL-EBI. http://www.ebi.ac.uk/training/online/glossary/term/509
2. Hayworth, Douglas. Overview of Protein-Protein Interaction Analysis. Thermo Scientific. http://www.piercenet.com/method/overview-protein-protein-interaction-analysis. Retrieved May 01, 2014.
3. Uniprot. http://www.uniprot.org/uniprot/P78527. Retrieved May 1, 2014.
4. Genetics Home Reference. (2010) http://ghr.nlm.nih.gov/gene/ERCC3. Retrieved May 1, 2014.
5. Croteau, D., Peng, Y., Van Houten, B. (2008) DNA repair gets physical: mapping a XPA binding site on ERCC1. DNA Repair. 7(5), 819-826. Available from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2494945/
6. Genetics Home Reference. (2010) http://ghr.nlm.nih.gov/gene/ERCC2. Retrieved May 1, 2014.
1. EMBL-EBI. http://www.ebi.ac.uk/training/online/glossary/term/509
2. Hayworth, Douglas. Overview of Protein-Protein Interaction Analysis. Thermo Scientific. http://www.piercenet.com/method/overview-protein-protein-interaction-analysis. Retrieved May 01, 2014.
3. Uniprot. http://www.uniprot.org/uniprot/P78527. Retrieved May 1, 2014.
4. Genetics Home Reference. (2010) http://ghr.nlm.nih.gov/gene/ERCC3. Retrieved May 1, 2014.
5. Croteau, D., Peng, Y., Van Houten, B. (2008) DNA repair gets physical: mapping a XPA binding site on ERCC1. DNA Repair. 7(5), 819-826. Available from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2494945/
6. Genetics Home Reference. (2010) http://ghr.nlm.nih.gov/gene/ERCC2. Retrieved May 1, 2014.
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison