Gene Expression Analysis
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
|
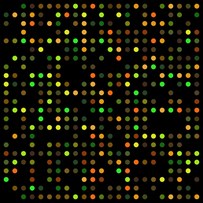
What is Gene Expression Analysis? Although every cell in the body contains the same genetic material, some genes are only expressed in a certain type of cell. [1] Knowing which genes are expressed (and which are not) in a specific type of cell can aid an understanding of how that cell should operate normally. Microarray technologies have been developed that allow researchers to observe the activity of thousands of genes at once. [1] Microarrays are created by attaching thousands of DNA sequences, or probes, to a microscope slide. These sequences are activated on the slide and begin to synthesize messenger RNA (mRNA), which is the precursor to proteins. These mRNAs are labeled with fluorescent tags, and the color of the tag and intensity of fluorescence can identify the genes that are being expressed. A brighter spot on the slide means that the particular gene is more active than others. [1] |
|
A public database has been created, called Gene Expression Omnibus (GEO),
which has collected array and sequence-based expression data to help
users query a wide variety of experiments and gene expression profiles. [2] These experiments cover a wide variety of cell types and experimental conditions, and can give insight to how a particular gene functions in different environments.
How is XPA represented in the GEO database?
XPA is well represented in the GEO database, but the studies that were performed exploited the function of XPA to be used in unrelated conditions. This means that although a search for "XPA" in the database will yield many results, there were not any studies that were specific to the disease, xeroderma pigmentosum. Despite the lack of specific studies to the disease, these experiments can be used to learn more about the way that XPA operates under a variety of conditions. An example of this application can be found below.
How is XPA represented in the GEO database?
XPA is well represented in the GEO database, but the studies that were performed exploited the function of XPA to be used in unrelated conditions. This means that although a search for "XPA" in the database will yield many results, there were not any studies that were specific to the disease, xeroderma pigmentosum. Despite the lack of specific studies to the disease, these experiments can be used to learn more about the way that XPA operates under a variety of conditions. An example of this application can be found below.
|
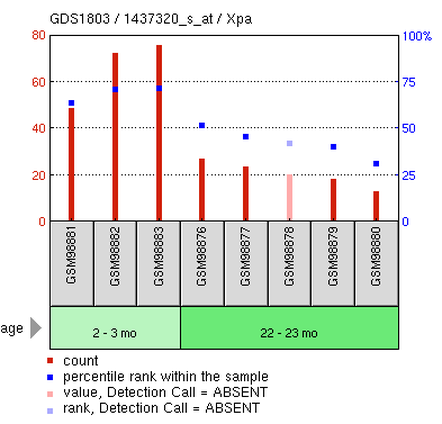
"Age Effect on Hematopoietic Stem Cells" Organism: Mus musculus How to interpret this chart:
Red bars are the count or value; quantification method specific to the study. Blue dots are the percentile rank within the sample. The pattern of these dots is important to understanding the overall trend that was observed. This chart shows higher expression in the younger samples and a decrease in expression in older samples. Grey bars are the individual samples that were used in the study. This particular study used eight different samples. Green bars show the conditions of the individual samples. These bars are perhaps the easiest way to interpret what the experiment was trying to study. In this particular study, the researchers observed the effect of age on XPA expression. They used two conditions: 2-3 month-old and 22-23 month-old stem cells. Results: This study suggests that younger stem cells show higher XPA expression than older stem cells. The expression profile can be accessed here. |
Analysis
This study, although interesting, is only tangentially related to my research on xeroderma pigmentosum. The experiment used stem cells to measure how XPA is expressed as a function of age, and suggests that XPA has lesser expression in older stem cells than younger stem cells. This may suggest decreased DNA repair in older individuals. This experiment would be an interesting starting point for future research in younger and older individuals and their ability to repair DNA with XPA in nucleotide excision repair.
This study, although interesting, is only tangentially related to my research on xeroderma pigmentosum. The experiment used stem cells to measure how XPA is expressed as a function of age, and suggests that XPA has lesser expression in older stem cells than younger stem cells. This may suggest decreased DNA repair in older individuals. This experiment would be an interesting starting point for future research in younger and older individuals and their ability to repair DNA with XPA in nucleotide excision repair.
References
1. DNA Microarray Technology. National Human Genome Research Institute. http://www.genome.gov/10000533. Retrieved May 05, 2014.
2. Gene Expression Omnibus. National Center for Biotechnology Information. http://www.ncbi.nlm.nih.gov/geo/. Retrieved April 29, 2014.
1. DNA Microarray Technology. National Human Genome Research Institute. http://www.genome.gov/10000533. Retrieved May 05, 2014.
2. Gene Expression Omnibus. National Center for Biotechnology Information. http://www.ncbi.nlm.nih.gov/geo/. Retrieved April 29, 2014.
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison