Protein Motifs
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What are motifs?
Sequence motifs are short and recurring patterns in DNA that are presumed to have some biological function. [1] Because sequence motifs often code for a specific process, motifs are highly conserved. Motifs can indicate sequence-specific binding sites and other important processes at the RNA level. [1] Motifs with known function can be a good indicator of the overall function gene or protein of interest.
|
Motifs in XPA
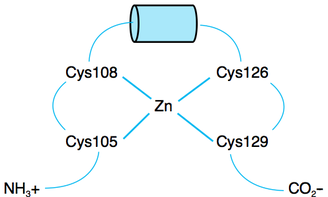
The literature describes a motif of utmost importance in the XPA protein, called a zinc finger motif. One of the main functions of zinc finger motifs is to bind DNA, and is often done in a sequence-specific manner. [2] Zinc finger motifs can coordinate one or more zinc atoms, and are extraordinarily versatile in their binding capabilities. [3] As concluded by Morita et al (1996), the zinc finger motif in XPA follows this pattern: Cys105-X2-Cys108-X17-Cys126-X2-Cys129
The literature describes the importance of the four cysteine residues that coordinate a zinc atom. If any of these cysteine residues are mutated, the protein is not capable of forming correctly, thus rendering it nonfunctional. [4] I was interested to see if there were any other motifs that helped contribute to the function of XPA.
|
Generating motifs of the XPA protein
Motifs were generated by submitting the protein sequence to a program called MEME (Multiple Em for Motif Elicitation). The database compares the given sequence to a motif database and generates a list of possible motifs in that sequence. [5]
Motifs were generated by submitting the protein sequence to a program called MEME (Multiple Em for Motif Elicitation). The database compares the given sequence to a motif database and generates a list of possible motifs in that sequence. [5]
How to read a MEME motif
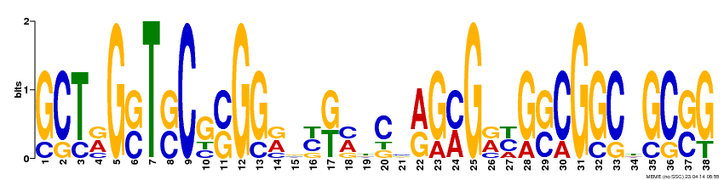
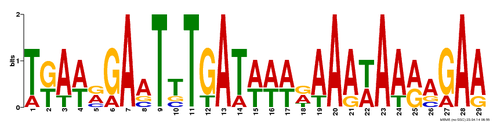
Motif predictions, pictured below, are difficult to interpret without a bit of background. The prediction of motifs is done by comparing how frequently a specific sequence recurs in the overall sequence, and these images are a visual representation of the probability that the particular nucleotide will be in that position. The number of nucleotides that are shown and their respective sizes indicate the likelihood of that nucleotide. For example, in Figure 1, the nucleotide at position 7 is very likely to be a thymine (T), whereas the nucleotide at position 4 could be guanine, adenine, or cytosine (G, A, C). In this second case (position 4), none of the three have a high probability relative to each other, or to the other nucleotides in the sequence.
Although it is important to understand how to read these figures, they still have little meaning. A program called GOMO can analyze the motif sequences that are generated by MEME, and assigns the most likely gene ontologies to the particular sequence. A score for each ontology was generated as the geometric mean of rank-sum test(s) to determine their significance. [6] The most relevant gene ontologies are reported below.
Motif predictions, pictured below, are difficult to interpret without a bit of background. The prediction of motifs is done by comparing how frequently a specific sequence recurs in the overall sequence, and these images are a visual representation of the probability that the particular nucleotide will be in that position. The number of nucleotides that are shown and their respective sizes indicate the likelihood of that nucleotide. For example, in Figure 1, the nucleotide at position 7 is very likely to be a thymine (T), whereas the nucleotide at position 4 could be guanine, adenine, or cytosine (G, A, C). In this second case (position 4), none of the three have a high probability relative to each other, or to the other nucleotides in the sequence.
Although it is important to understand how to read these figures, they still have little meaning. A program called GOMO can analyze the motif sequences that are generated by MEME, and assigns the most likely gene ontologies to the particular sequence. A score for each ontology was generated as the geometric mean of rank-sum test(s) to determine their significance. [6] The most relevant gene ontologies are reported below.
|
Gene Ontology
Cellular Component: transcription factor complex GOMO score: 1.512e-05 Molecular Function: zinc ion binding GOMO score: 2.931e-04 Biological Process: G-protein coupled receptor protein signaling pathway GOMO score: 2.004e-04 Biological Process: G-protein coupled receptor protein signaling pathway GOMO score: 1.415e-04 Biological Process: DNA damage checkpoint GOMO score: 5.022e-03 |
Discussion
All three motifs shared at least one similar biological process or molecular function with XPA. Due to discrepancies in the sequence, many more possible ontologies were generated, but they were not relevant (760 ontologies for Motif #1, 20 for Motif #2, and 21 for Motif #3). All of the above ontologies were both statistically significant and had a high percent specificity. The most intriguing ontologies include the zinc ion binding molecular function in Motif #1 and the DNA damage checkpoint biological process in Motif #3. The presence of zinc ion binding in Motif #1 suggests a correlation between Motif #1 and the zinc finger motif in the literature. This correlation should be further investigated to determine if the motif generated by MEME is a part of the motif from the literature, or if they are two separate motifs. The biological process of DNA damage checkpoint in Motif #3 is also interesting because XPA binds to damaged DNA in the nucleotide excision repair process. This motif may suggest a biological process that allows XPA to recognize if the DNA is damaged, and should also be investigated further.
References
1. D'haeseleer, Patrik. What are DNA sequence motifs? Nature Biotechnology. 2010. 24, 423 - 425. Available from http://www.ncbi.nlm.nih.gov/pubmed/16601727.
2. Jennifer McDowall. Zinc Fingers. InterPro. http://www.ebi.ac.uk/interpro/potm/2007_3/Page2.htm. Retrieved May 07, 2014.
3. Jennifer McDowall. Zinc Fingers. InterPro. http://www.ebi.ac.uk/interpro/potm/2007_3/Page1.htm. Retrieved May 07, 2014.
4. Morita, et al. (1996) Implications of the zinc-finger motif found in the DNA-binding domain of the human XPA protein. Genes to Cells. 1, 437–442. Available from http://www.ncbi.nlm.nih.gov/pubmed/9078375.
5. Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
6. Fabian A. Buske, Mikael Boden, Denis C. Bauer and Timothy L. Bailey, "Assigning roles to DNA regulatory motifs using comparative genomics", Bioinformatics, 26(7), 860-866, 2010.
1. D'haeseleer, Patrik. What are DNA sequence motifs? Nature Biotechnology. 2010. 24, 423 - 425. Available from http://www.ncbi.nlm.nih.gov/pubmed/16601727.
2. Jennifer McDowall. Zinc Fingers. InterPro. http://www.ebi.ac.uk/interpro/potm/2007_3/Page2.htm. Retrieved May 07, 2014.
3. Jennifer McDowall. Zinc Fingers. InterPro. http://www.ebi.ac.uk/interpro/potm/2007_3/Page1.htm. Retrieved May 07, 2014.
4. Morita, et al. (1996) Implications of the zinc-finger motif found in the DNA-binding domain of the human XPA protein. Genes to Cells. 1, 437–442. Available from http://www.ncbi.nlm.nih.gov/pubmed/9078375.
5. Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
6. Fabian A. Buske, Mikael Boden, Denis C. Bauer and Timothy L. Bailey, "Assigning roles to DNA regulatory motifs using comparative genomics", Bioinformatics, 26(7), 860-866, 2010.
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison