Future Research
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
|
The slides from my final presentation can be viewed at the right. |
| ||||||
Conclusions
Although xeroderma pigmentosum is rare in the United States, it is certainly a public health issue of great concern. Patients with xeroderma pigmentosum are at high risk for skin cancer; research shows about fifty percent of patients will have their first skin cancer by age ten. [1] Xeroderma pigmentosum inhibits the mechanism that aids in repairing DNA damage, and thus, leaves damaged DNA unattended. Since there is no cure for xeroderma pigmentosum, any and all preventive measures should be taken to protect DNA from UV-induced damage to prevent the onset of skin cancer.
Although xeroderma pigmentosum is rare in the United States, it is certainly a public health issue of great concern. Patients with xeroderma pigmentosum are at high risk for skin cancer; research shows about fifty percent of patients will have their first skin cancer by age ten. [1] Xeroderma pigmentosum inhibits the mechanism that aids in repairing DNA damage, and thus, leaves damaged DNA unattended. Since there is no cure for xeroderma pigmentosum, any and all preventive measures should be taken to protect DNA from UV-induced damage to prevent the onset of skin cancer.
|
Some preventive measures for cancer in unaffected individuals include lifestyle adaptations, such as a well balanced diet. Some foods have natural chemicals that support biological processes and pathways in the body, and thus, those foods are recommended to be eaten more frequently. Broccoli and other cruciferous vegetables are highly recommended because it contains the chemical sulforaphane, which is said to help reduce cancer risk. [2] In the cell, sulforaphane is normally active in an oxidative stress pathway; this pathway is another method used to reduce precursors for cancer. In this pathway, sulforaphane acts in an intermediate step that involves releasing zinc from a dominant protein in the pathway. This step is vital to the completion of the pathway, so it can be inferred that an increase in sulforaphane in the cell would increase the efficiency of this step.
|
It is still unclear if sulforaphane is able to release zinc from other pathways in the cell. Zinc is a key component in many biological molecules, including the xeroderma pigmentosum protein, XPA. The XPA protein contains a zinc finger domain that facilitates binding to damaged DNA, and is composed of similar elements as the aforementioned oxidative stress protein. Recent studies have shown that targeting XPA with sulforaphane can decrease repair efficiency, and have proposed that sulforaphane is able to remove zinc from XPA in the same manner as the oxidative stress pathway. [3] Without zinc, the zinc finger domain cannot bind damaged DNA, and this will render the protein nonfunctional. Without a properly functioning XPA protein, the nucleotide excision repair process is severely inhibited, and DNA cannot be repaired.
The primary goal of my research is to identify how sulforaphane affects DNA repair machinery. Using genomic and proteomic techniques, I will perform the following three experiments to test my hypotheses that sulforaphane can inhibit the primary binding function of XPA. I will perform my studies in a mouse model because of its high homology and similar phenotype to humans.
|
Aim 1:
To determine if sulforaphane eliminates binding between XPA and ERCC1. XPA has many binding partners, but one of the most important binding partners is the protein ERCC1. ERCC1 is responsible for forming a dual protein complex with XPF, and this complex allows nucleotide excision repair to be completed. [4] The role of XPA in the cell is to recruit ERCC1 so that this larger complex can be formed. XPA is able to recognize that DNA is damaged in the cell, and this recognition sends a signal to ERCC1 and other proteins to be recruited. [4] However, if sulforaphane is able to inhibit the binding abilities of XPA, XPA may not be able to bind to damaged DNA and other proteins. If XPA cannot bind, the cycle of DNA repair will be inhibited.
|
|
|
I will perform a yeast-2-hybrid assay in order to test the binding abilities of XPA and ERCC1 with and without sulforaphane. I am using a mouse model for its phenotypic and proteomic similarity, so to maintain consistency, I will insert mouse DNA onto a plasmid, and insert that plasmid into yeast.
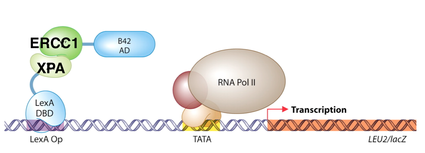
First, some quick background (or review): the yeast-2-hybrid assay is an experiment that tests the binding abilities of two proteins by exploiting a natural process in yeast. To turn on a gene, a special protein complex must bind to the operator of that particular gene (called LexA Op in Figures 2 and 3, for example.) Within this special protein are two distinct units, called the Activation Domain and the DNA Binding Domain. The DNA binding domain allows the complex to bind to the operator, and the activation domain allows the gene to be activated, or turned on. As long as these two domains are able to bind and communicate, the gene will be turned on. For this reason, we are able to attach proteins to these domains and test their binding abilities. If the proteins bind, the domains will be able to communicate, and transcription of the gene will be turned on. If the proteins do not bind, however, the activation domain will not be able to communicate with the DNA Binding Domain, and transcription of the gene will not be possible. |
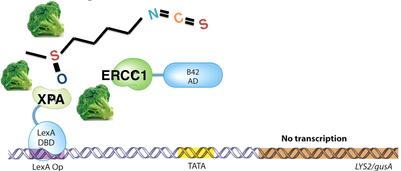
I expect that my results will be proof of principle that sulforaphane is able to inhibit binding between XPA and ERCC1. I expect that when XPA and ERCC1 are exposed to each other without sulforaphane, they will be able to bind, and subsequent transcription will be observed (Figure 2). However, I expect that when XPA and ERCC1 are exposed to sulforaphane, binding will be inhibited, and transcription will not occur (Figure 3).
|
Aim 2:
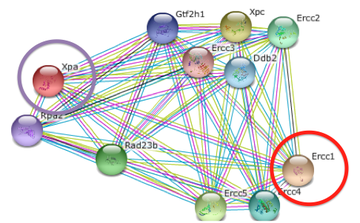
To identify the transcripts that show decreased expression as a result of sulforaphane. I am also interested in studying how a lack of binding by XPA can affect the other proteins in the cell. In the STRING database, I organized XPA's binding partners based on the order in which they bind in nucleotide excision repair. This was a convenient way to organize the associated proteins, because nucleotide excision repair protein binding occurs in a very specific order and binding is dependent on signals from other proteins. [4] RNA seq is a method used to measure the abundance of transcripts of different genes under different conditions. This method is an efficient way to observe how a genetic or environmental condition can affect the way that transcripts are produced, which will lead to the translation into protein. I will use four experimental conditions in my RNA seq experiment: wild type, wild type exposed to sulforaphane, mutant, and mutant exposed to sulforaphane. I expect to see two distinct patterns from RNA seq in all of the conditions. I think that the transcripts that would normally be expressed after XPA binds to DNA will have decreased expression. I expect this result because XPA will not be present to recruit the proteins to the site of damage, so there will be no signal for the transcripts to be produced. On the other hand, I expect to see an increase in transcripts for proteins that bind prior to XPA. I think that the transcripts will be overproduced because there will not be a signal from XPA that the process is still working properly. Overall, I think that the most severe effects will be observed when there is both a mutation in XPA and sulforaphane is present. I expect that the mutation and sulforaphane will affect the transcripts twice as much, because it is inhibiting the protein in two different ways. A similar pattern will be seen with either the mutation OR presence of sulforaphane, but it will not be as severe. |
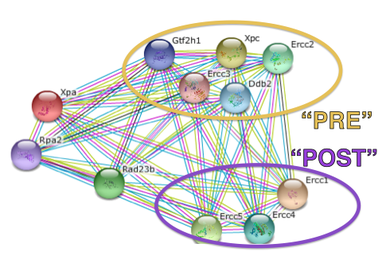
Figure 4: STRING figure of all binding partners of XPA. Proteins are divided into two categories based on their binding order in nucleotide excision repair. Proteins that bind before XPA are annotated as "PRE" and are circled in yellow, proteins that bind after XPA are annotated as "POST" and are circled in purple.
|
Aim 3:
To quantify the change in expression of XPA and other DNA repair proteins in the presence of sulforaphane over time.
To quantify the change in expression of XPA and other DNA repair proteins in the presence of sulforaphane over time.
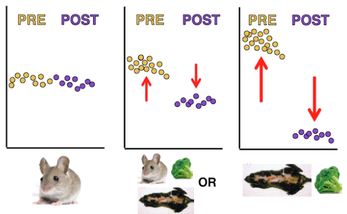
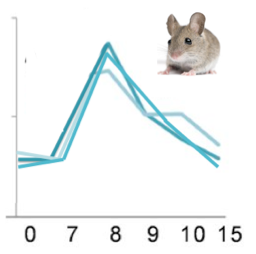
Finally, I was interested in seeing how protein expression over time would be affected by the same conditions from Aim 2: the presence of a mutation, sulforaphane, or both. Using a quantiative proteomic method, Figure 6 shows my initial prediction for protein expression. I expect that DNA repair proteins will be expressed at relatively low levels in the cell when there is no damage to be repaired. However, when the cell recognizes that there is damaged DNA to repair, I think that the proteomic response will be intense and immediate. As the damage is repaired, the proteomic expression will decrease back to the original levels.
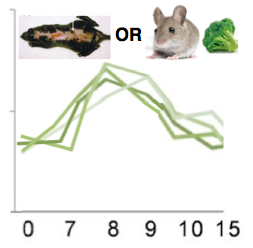
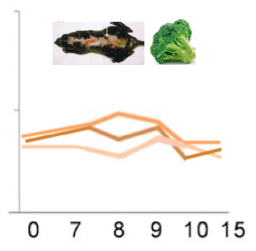
I expect to see differences arise in proteomic expression when sulforaphane is present, XPA is mutated, or both. I think that a mouse with a mutation or a wild type mouse exposed sulforaphane will show a similar pattern of expression; the response time will be slower, and the abundance will be lower. This would mean that the proteins are not working as they are supposed to, but they are still somewhat functional. Conversely, I think that a mutant mouse that is exposed to sulforaphane would have considerably decreased expression (Figure 8). I think that the combination of the mutation and exposure to sulforaphane would decrease the ability of the proteins to function, and thus, low expression would be observed via quantitative proteomics.
I expect to see differences arise in proteomic expression when sulforaphane is present, XPA is mutated, or both. I think that a mouse with a mutation or a wild type mouse exposed sulforaphane will show a similar pattern of expression; the response time will be slower, and the abundance will be lower. This would mean that the proteins are not working as they are supposed to, but they are still somewhat functional. Conversely, I think that a mutant mouse that is exposed to sulforaphane would have considerably decreased expression (Figure 8). I think that the combination of the mutation and exposure to sulforaphane would decrease the ability of the proteins to function, and thus, low expression would be observed via quantitative proteomics.
Discussion
If my hypothesis is correct and sulforaphane does inhibit the ability of XPA to bind, I think that this could have significant implications for individuals with xeroderma pigmentosum. Namely, patients with xeroderma pigmentosum would be advised to remove broccoli and other vegetables containing sulforaphane from their diet. However, I think that this recommendation would not carry as much weight as medicine or other therapies; I believe the response would be similar to when unaffected individuals are told to eat broccoli for its chemopreventive properties, which only has a moderate response rate. While an increase of broccoli in the diet can benefit an individual, it is not likely be the determining factor of avoiding cancer. Likewise, I don't think that eliminating broccoli from the diet of an individual with xeroderma pigmentosum will cure their inability to repair damaged DNA. However, I think that however small the benefit, any preventive measures should be taken. In the future, this research could lead to the discovery of effects of other natural chemicals in foods and their effects on xeroderma pigmentosum and other diseases.
References
1. Friedberg, E. (2001). How Nucleotide Excision Repair Protects Against Cancer. Nature Reviews, 1, 22-33. Available from http://classes.biology.ucsd.edu/old.web.classes/bggn220.FA03/Friedberg%20rev%20DNA-NER.pdf.
2. Gupta, P., Kim, B., Kim SH., Srivastava, SK. (2014, February 10) Molecular targets of isothiocyanates in cancer: Recent advances. Molecular Nutrition and Food Research 2014: 00, 1–23. doi: 10.1002/mnfr.201300684. Available from http://www.ncbi.nlm.nih.gov/pubmed/24510468.
3. Piberger AL., Köberle B., Hartwig A. The broccoli-born isothiocyanate sulforaphane impairs nucleotide excision repair: XPA as one potential target. (2013, December 19) Molecular Toxicology 2014: 88(3): 647-58. doi: 10.1007/s00204-013-1178-2. Available from http://www.ncbi.nlm.nih.gov/pubmed/?term=xpa%20sulforaphane.
4. Croteau, D., Peng, Y., & Van Houten, B. (2008). DNA repair gets physical: mapping a XPA binding site on ERCC1. DNA Repair, 7(5), 819-826. Available from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2494945/
1. Friedberg, E. (2001). How Nucleotide Excision Repair Protects Against Cancer. Nature Reviews, 1, 22-33. Available from http://classes.biology.ucsd.edu/old.web.classes/bggn220.FA03/Friedberg%20rev%20DNA-NER.pdf.
2. Gupta, P., Kim, B., Kim SH., Srivastava, SK. (2014, February 10) Molecular targets of isothiocyanates in cancer: Recent advances. Molecular Nutrition and Food Research 2014: 00, 1–23. doi: 10.1002/mnfr.201300684. Available from http://www.ncbi.nlm.nih.gov/pubmed/24510468.
3. Piberger AL., Köberle B., Hartwig A. The broccoli-born isothiocyanate sulforaphane impairs nucleotide excision repair: XPA as one potential target. (2013, December 19) Molecular Toxicology 2014: 88(3): 647-58. doi: 10.1007/s00204-013-1178-2. Available from http://www.ncbi.nlm.nih.gov/pubmed/?term=xpa%20sulforaphane.
4. Croteau, D., Peng, Y., & Van Houten, B. (2008). DNA repair gets physical: mapping a XPA binding site on ERCC1. DNA Repair, 7(5), 819-826. Available from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2494945/
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison