Protein Homology
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What is homology?
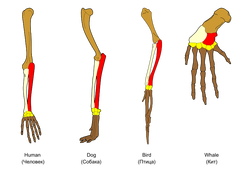
Homologous characters are those that are descended from a common ancestor. [1] A common example of a homologous structure can be seen in the picture at the right, which shows a similar limb structure that is found in humans, dogs, birds, and whales. All four structures came from a common ancestor, and their differences are due to modification over evolutionary time.
The study of homology can also be applied to smaller structures, such as proteins. Similar to the limb described above, proteins can be modified over evolutionary time. By studying the same protein in different organisms, we can see how the protein was modified over time, based on the differences we see in other organisms. In addition, the presence of a homolog in a model organism makes it possible to study the protein in more depth.
Homologs of the XPA gene
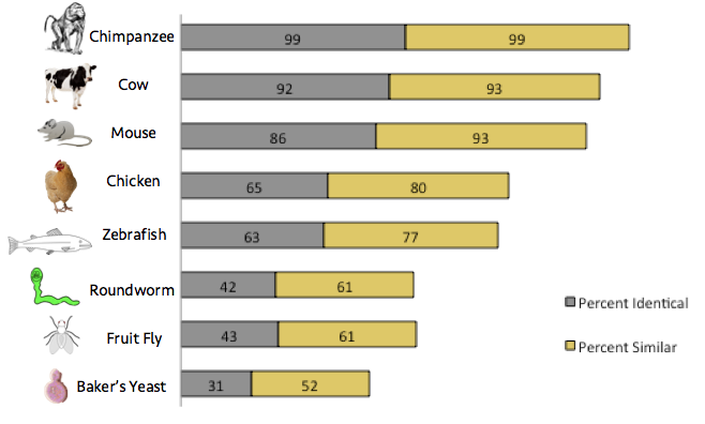
Homology comparisons were found using NCBI's HomoloGene and BLAST (Basic Local Alignment Search Tool). HomoloGene provides information about the known homologs of the gene and protein in a number of organisms, including model organisms. This list is a useful starting point for further investigation, and the HomoloGene results for the XPA gene can be found here. To further the research, BLAST was used to find the amount of similarity and identity to the human protein in each homolog. BLAST finds regions of similarity between sequences and calculates the statistical significance of those matches. [2] The results of the BLAST comparison can be found in the figure below.
Discussion
The results of this comparison show that the highest amount of similarity can be found in the chimpanzee, cow, and mouse, and the least amount of similarity is found in the roundworm, fruit fly, and baker's yeast. Xeroderma pigmentosum is a mammalian disease, and the most common effects of XP are related to the skin. Since the chimpanzee, cow, and mouse are evolutionarily the most similar to humans, it makes sense that that would be most related. It is not surprising that the roundworm, fruit fly, and baker's yeast have less than 50% identity, because they are evolutionarily quite different organisms to humans. In addition, the yeast homolog is the only protein of all of the tested organisms that does not include "xeroderma pigmentosum" in its name, which suggests that the homolog serves a similar but not identical function. A low E value was found for all organisms, which shows that all tests are statistically significant.

Human (Homo sapiens)
DNA repair protein complementing XP-A cells Accession Number: NP_000371.1 Protein Sequence: FASTA 
Chimpanzee (Pan troglodytes)
Predicted: DNA repair protein complementing XP-A cells Accession Number: XP_001156167.2 Identical: 99% // Similar: 99% E value: 0.0 Protein sequence: FASTA 
Cow (Bos taurus)
DNA repair protein complementing XP-A cells Accession Number: NP_001033770.1 Identical: 92% // Similar: 93% E value: 3e-170 Protein sequence: FASTA 
Mouse (Mus musculus)
DNA repair protein complementing XP-A cells homolog Accession Number: NP_035858.2 Identical: 86% // Similar: 93% E value: 1e-150 Protein sequence: FASTA 
Chicken (Gallus gallus)
DNA repair protein complementing XP-A cells homolog Accession Number: NP_990184.1 Identical: 65% // Similar: 80% E value: 9e-119 Protein sequence: FASTA |

Zebrafish (Danio rerio)
Xeroderma pigmentosum, complementation group A Accession Number: NP_956765.1 Identical: 63% // Similar: 77% E value: 5e-98 Protein sequence: FASTA 
Roundworm (Caenorhabditis elegans)
Protein XPA-1 Accession Number: NP_492025.1 Identical: 42% // Similar: 61% E value: 4e-48 Protein sequence: FASTA 
Fruit Fly (Drosophila melanogaster)
Xeroderma pigmentosum group A-like Accession Number: NP_476866.1 Identical: 43% // Similar: 61% E value: 5e-72 Protein sequence: FASTA 
Baker's Yeast (Saccharomyces cerevisiae)
Rad14p Accession Number: NP_013928.1 Identical: 31% // Similar: 52% E value: 7e-12 Protein sequence: FASTA |
References
1. Brinkmann H, Delsuc Frederic, Philippe H. Phylogenomics and the Reconstruction of the Tree of Life. Nature Review, 2005;6:361-375. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15861208 PubMed PMID: 15861208
2. BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi. Accessed February 18, 2014.
From figure (yeast image): http://www.sciencedaily.com/releases/2007/05/070530102851.htm
1. Brinkmann H, Delsuc Frederic, Philippe H. Phylogenomics and the Reconstruction of the Tree of Life. Nature Review, 2005;6:361-375. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15861208 PubMed PMID: 15861208
2. BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi. Accessed February 18, 2014.
From figure (yeast image): http://www.sciencedaily.com/releases/2007/05/070530102851.htm
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison