Mutant Phenotypes & RNAi
This page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Mutants and Model Organisms
Experiments on the genomic level often cannot be performed in humans due to ethical reasons. As a result, well-studied organisms with high homology and similar biological processes to humans are used as a replacement. Common model organisms include mice, fruit flies, roundworms, and yeast. An organism can be defined as a "model organism" by many criteria. Criteria of great importance is that the phenotype of the gene being studied is easy to detect in the organism, and that it exhibits a similar phenotype to a human model. Other criteria of note are that the organism is easy to care for, has a short generation time, is inexpensive, and is relatively simple to work with. [1] With the help of model organisms, research can be performed on a specific gene in an ethical, cost-effective, and time-effective manner. Most of the research of human disorders originated in one of the aforementioned organisms before it was tested on a human scale.
What is RNAi?
RNA Interference, or RNAi, is a way of disrupting a gene to mimic a mutation without altering the genetic material. RNAi is a natural process in the body, but in a research setting, the mechanism can be exploited to target specific genes. Through a complex mechanism, RNAi is able to destroy molecular messengers that code for protein expression. Without these messengers, the gene's function is "turned off" because the corresponding protein cannot be made. [2] RNAi is an important avenue for future research, because researchers can shut off individual genes, one at a time, to see how various processes are affected. This knowledge can lead toward development of drugs or treatment options for a variety of diseases. [2]
|
Mutant Phenotypes Many databases were used to search for mutant phenotypes across model organisms, including Saccharomyces Genome Database, The Zebrafish Model Organism Database, Phenobank/Wormbase, FlyBase, and Mouse Genome Informatics. These databases were used because the associated organisms have known homologs of the XPA gene. RNAi was used to cause a mutation in many of these organisms. The Saccharomyces Genome Database was used to find the mutant phenotype in yeast. Without Rad14, the yeast homologous gene, yeast showed decreased UV resistance. [3] The Zebrafish Model Organism Database (ZFIN) generated inconclusive data for the phenotype in zebrafish. ZFIN reports that zebrafish have the phenotype of the human ortholog, but it is not clear how the phenotype expresses itself specifically in zebrafish. [4] According to Phenobank and Wormbase, the XPA homolog (xpa-1) in C. elegans is needed for meiosis, germ cell apoptosis, and survival of germ and somatic cells, and the gene can be detected throughout the entire life cycle. [5] Loss of activity in xpa-1 results in hypersensitivity to UV irradiation at all stages of development. [6] FlyBase reports that Drosophila (fruit fly) with a nonfunctional XPA gene are viable, but no phenotypic data is available. [7]
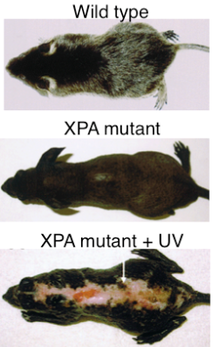
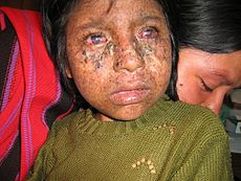
Finally, Mouse Genome Informatics reports that when exposed to UV light, mouse homozygous null mutants are highly susceptible to tumors associated with skin, eyes, lungs, tongue, and liver. [8] Mice exhibit the most similar phenotype to humans; a comparison can be seen in Figures 1 and 2. |
Discussion
After analyzing the results, it became apparent that mice are the obvious choice for studying xeroderma pigmentosum. Although they are the most complex organism of the four, they have an extraordinarily similar phenotype to humans, which suggests that similar mechanisms are shared between the two organisms. Despite the cost and longer life cycle, it is important to study a disease in an organism that shares a similar phenotype and genetic mechanism. Thus, mice would be the ideal organism to study this disease.
After analyzing the results, it became apparent that mice are the obvious choice for studying xeroderma pigmentosum. Although they are the most complex organism of the four, they have an extraordinarily similar phenotype to humans, which suggests that similar mechanisms are shared between the two organisms. Despite the cost and longer life cycle, it is important to study a disease in an organism that shares a similar phenotype and genetic mechanism. Thus, mice would be the ideal organism to study this disease.
References
1. Flanagan, Emma. Fastbleep Biology Notes. http://www.fastbleep.com/biology-notes/33/110/815. Retrieved May 16, 2014.
2. The RNAi Center. RNAi at a Glance. http://www.liai.org/pages/rnai-at-a-glance. Retrieved May 16, 2014.
3. Saccharomyces Genome Database. http://www.yeastgenome.org/cgi-bin/locus.fpl?locus=rad14. Retrieved May 16, 2014.
4. The Zebrafish Model Organism Database. http://zfin.org/ZDB-GENE-040426-1205. Retrieved May 16, 2014.
5. Wormbase. http://www.wormbase.org/species/c_elegans/gene/WBGene00006963?query=xpa-1#0b-9e-3. Retrieved May 16, 2014.
6. Phenobank. http://www.worm.mpi-cbg.de/phenobank/cgi-bin/GenePage.py?GeneID=510968. Retrieved May 16, 2014.
7. FlyBase. http://flybase.org/reports/FBgn0004832.html. Retrieved May 16, 2014.
8. Mouse Genome Informatics. http://www.informatics.jax.org/marker/MGI:99135. Retrieved May 16, 2014.
9. Yamazaki, F., Okamoto, H., Matsumura, Y., Tanaka, K., Kunisada, T., & Horio, T. (2005) Development of a New Mouse Model (Xeroderma Pigmentosum A-Deficient, Stem Cell Factor-Transgenic) of Ultraviolet B-Induced Melanoma. Journal of Investigative Dermatology, 125, 521–525. doi:10.1111/j.0022-202X.2005.23753.x. Available from http://www.nature.com/jid/journal/v125/n3/full/5603519a.html.
1. Flanagan, Emma. Fastbleep Biology Notes. http://www.fastbleep.com/biology-notes/33/110/815. Retrieved May 16, 2014.
2. The RNAi Center. RNAi at a Glance. http://www.liai.org/pages/rnai-at-a-glance. Retrieved May 16, 2014.
3. Saccharomyces Genome Database. http://www.yeastgenome.org/cgi-bin/locus.fpl?locus=rad14. Retrieved May 16, 2014.
4. The Zebrafish Model Organism Database. http://zfin.org/ZDB-GENE-040426-1205. Retrieved May 16, 2014.
5. Wormbase. http://www.wormbase.org/species/c_elegans/gene/WBGene00006963?query=xpa-1#0b-9e-3. Retrieved May 16, 2014.
6. Phenobank. http://www.worm.mpi-cbg.de/phenobank/cgi-bin/GenePage.py?GeneID=510968. Retrieved May 16, 2014.
7. FlyBase. http://flybase.org/reports/FBgn0004832.html. Retrieved May 16, 2014.
8. Mouse Genome Informatics. http://www.informatics.jax.org/marker/MGI:99135. Retrieved May 16, 2014.
9. Yamazaki, F., Okamoto, H., Matsumura, Y., Tanaka, K., Kunisada, T., & Horio, T. (2005) Development of a New Mouse Model (Xeroderma Pigmentosum A-Deficient, Stem Cell Factor-Transgenic) of Ultraviolet B-Induced Melanoma. Journal of Investigative Dermatology, 125, 521–525. doi:10.1111/j.0022-202X.2005.23753.x. Available from http://www.nature.com/jid/journal/v125/n3/full/5603519a.html.
Site Created By: Sarah Drewes
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison
Contact: [email protected]
Last Modified: 05/18/14
University of Wisconsin-Madison